Research
Evolution can happen in a relatively short time. Two best examples of rapid evolution in plants are crop domestication and weed adaptation. Therefore, I investigate the genomic changes in these plants over evolutionary time, and ask how genomic change influencing on evolutionary process. Utilizing vast amount of publicly available and new generated NGS data and developed bioinformatics tools, my research leverage approaches in population genetics, comparative genomics, forward and reverse genetics and biodiversity informatics to study diverse topics in evolutionary biology.
My research special focus on crop and weedy plant in Brassica and rice, major questions are (1) evolutionary history of rapid evolve species; (2) genomic changes associated with rapid evolutionary process; (3) genetic architecture of adaptive traits in fast evolve species; (4) genetic responses to ecological changes of abiotic and biotic factors. The ultimate goal is applying evolutionary genomics information to agricultural production improvement, weed/invasive species control and endangered species conservation so that human beings have a more sustainable living environment.
My research special focus on crop and weedy plant in Brassica and rice, major questions are (1) evolutionary history of rapid evolve species; (2) genomic changes associated with rapid evolutionary process; (3) genetic architecture of adaptive traits in fast evolve species; (4) genetic responses to ecological changes of abiotic and biotic factors. The ultimate goal is applying evolutionary genomics information to agricultural production improvement, weed/invasive species control and endangered species conservation so that human beings have a more sustainable living environment.
Brassica domesticationPlant domestication is the process by which wild plants evolve into a new form of crop plant through artificial selection to meet human needs (Doebley et al., 2006). Fundamental questions in plant domestication include where, when, how many times, and why it happened, and what kind of morphological and genetic changes are associated with this domestication process (Zeder 2015).
My research on plant domestication currently focuses on Brassica, a genus which includes many crops such as turnip, pak choi, Chinese cabbage, oilseed, broccoli, cauliflower, kale, cabbage, brussel sprouts and so on. All of these Brassica crops belong to three closely related species: Brassica rapa (2n =20), Brassica oleracea (2n =18) and the allopolyploid Brasscia napus (2n = 38). My current research project with Dr. Michael Barker and Dr. Chris Pires has been generating hundreds of transcriptomes from worldwide Brassica crop collections using Next Generation Sequencing platform. By analyzing these data, we hope to understand the domestication history of major Brassica crops, and the genetic changes associated with their domestication. We also hope to uncover how the genome triplication event shared among all Brassica lineages contributes to the diversification and morphological variation of these crops. Together, we hope these information will promote our understanding of plant domestication processes, and shed light on the future of molecular breeding. |
* "Triangle of U" diagram shown the genetic relationships between six species of the genus Brassica.
|
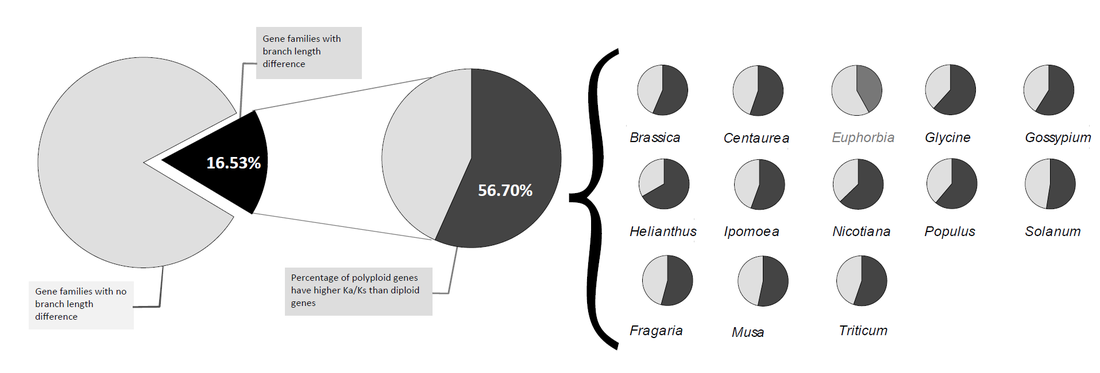
Comparisons of the substitution rate and deleterious mutations in polyploid and diploid plantsGenome duplication due to polyploidy has a profound impact on plants phenotypically and genetically. Here, we utilized pre-existing transcriptome data to test whether polyploidy in general has (1) under relaxed purifying selection and (2) accumulate more deleterious mutation when comparing with its diploid sister species. Our results indicate polyploid lineages were under relaxed purifying selection and accumulated more deleterious mutations than diploid plants. The faster accumulation of deleterious mutations in polyploids might be a major constraint to the long-term survival of a polyploid lineage after its establishment.
|
* Percentage of polyploidy copies have a higher Ka/Ks ratio than diploid in all gene families shown significant LRT test. Euphorbia is the only genus has less than 50%. 【Manuscript in preparation】.
|
Weedy rice: de-domestication and parallel evolution
|
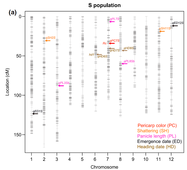
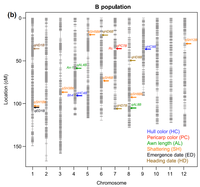
* QTL locations of Straw hull (S) and Black hull awned (B) US weedy strains.
|
Weedy rice is a conspecific weedy relative of rice which infests rice fields worldwide. These weedy strains originated multiple times independently from different rice cultivars. My postdoc research with Dr. Kenneth Olsen at Washington University in St. Louis focused on (1) the genetic basis of weedy traits in two major US weedy rice accessions, and (2) comparing the weedy rice origin in regions with and without wild rice co-occurred to uncover the role of wild rice during weedy rice origin.
For the first aim, we performed comparative QTL mapping of weediness traits using two recombinant inbred line populations derived from crosses between an indica cultivar and each of the two independently evolved US weedy rice strains. For most quantitative traits, we found little evidence of shared genetic bases between the two mapping populations, which suggests different underlying genetic mechanisms of weed parallel evolution. More details about this research is available here. For the second aim, we used 58708 SNPs generated using Genotyping-by-sequencing methods to examine the population structure of 224 rice collections, including weedy, cultivar and wild rice from five Southeast Asian countries, and weedy form from South Korea and USA. Our results suggest: (1) in regions wild rice co-occurred with cultivar rice, weedy rice is more prone to origin from wild rice introgression to crop than crop de-domestication; (2) De-domesticated weedy rice is commonly found in regions without wild rice. (3) Indica background weedy rice is more common than japonica and aus type in the regions studied (4) weedy rice can evolve very fast by gene introgression between cultivars and with wild rice.【Manuscript in preparation】. |
General phylogeographic pattern in East Asia, specially in CJK region
East Asia has the most diverse temperate flora of the world, and was the most important glacial refuge during the Quaternary ice age (Qiu et al., 2012). In contrast with Europe and North America, East Asia was not covered by unified large ice sheet during the last glacial maximum. Therefore, the phylogeographic patterns in this region are more complex than analogous regions in Europe and North America.
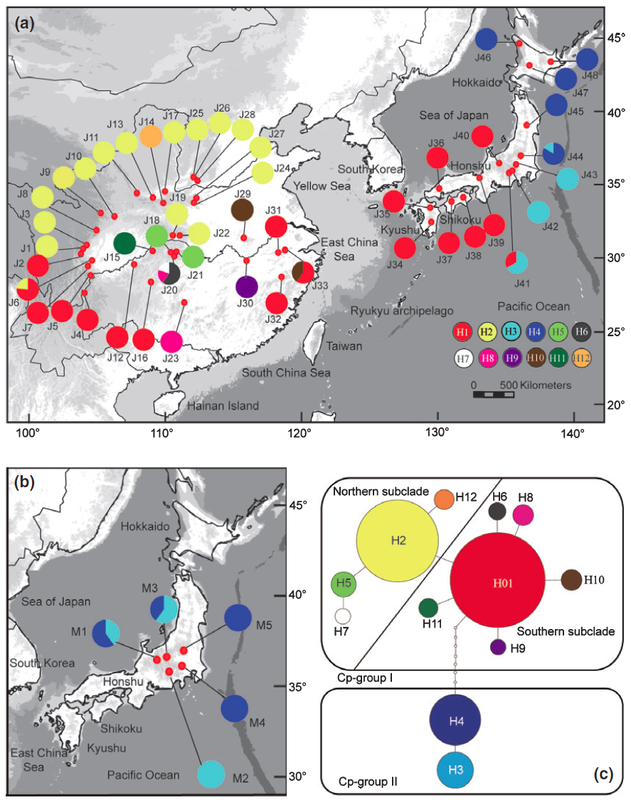
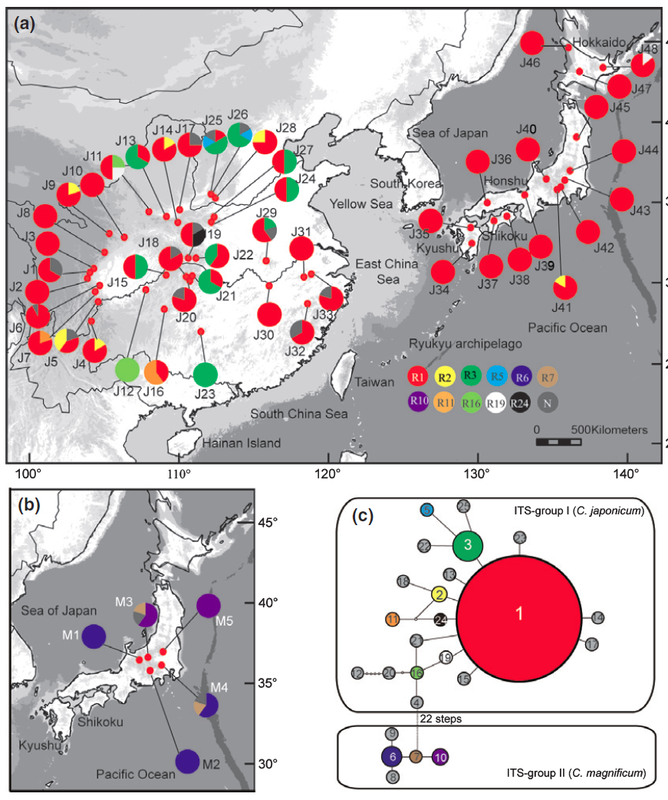
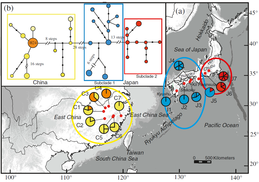
In the CJK region (east China, Korea and Japan), the fluctuations of sea level during the Quaternary climate change likely created opportunities for species migration and diversification in and around the exposed East China Sea. My Ph.D. study, with Dr. Yingxiong Qiu at Zhejiang University, focused on two rare endemic plant genera, Platycrater and Cercidiphyllum in this CJK region, the phylogeographic study of the two genera provide case studies for us to understanding the plant demographic history in this region. Cercidiphyllum Cercidiphyllum is the only genus in Cercidiphyllaceae, and there are only two extant species: C. japonicum and C. magnificum. The former one widely distributed in temperate regions in China and Japan, while the later one only restricted in central Japan. We used multiple molecular markers (cpDNA, ITS and SSR) in conjunction with Ecological Niche Modeling (ENM) to clarify the evolutionary relationship of the two sister species, and their demographic history. Our analysis results suggest C. japonicum invaded from East China to Japan via East China Sea land bridge during glacial period, the gene exchange event with C. magnificum in central Japan resulting the C. magnificum chloroplast genome introgressed to C. japonicum. The following northward population expansion during postglacial leads to the fixation of the introgressed chlorotypes. More details about this research is available here. Platycrater arguta Platycrater arguta (Hydrangeaceae) is a shrub that exhibits a disjunctive distribution in East China (var. sinensis) and South Japan (var. arguta), which provides us a great example to study the migration and gene exchange across the East China Sea region, especially testing the role of the land bridge between East China and South Japan during the last glacial maximum. Combined microsatellites, nuclear and chloroplast marker, our results show the two lineages diverged before the last glacial maximum, and no evidence of gene flow between the two varieties since then. Together with case studies of other species, like Cercidiphyllum, our results show a strong “filter” effect of the East China Sea for temperate plant species in East Asia. More details about this research is available here and here. |
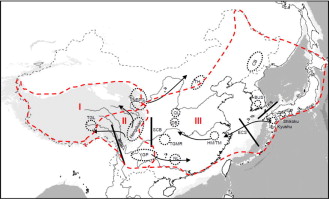
*Differing from Europe and North America, East Asia was not largely covered by ice sheet during the Last Glacial Maximum (18000 years ago).
*Summary of the phylogeographic patterns in East Asia. From Qiu et al., 2011
*Distribution of chloroplast (left) and ITS (right) genotype of C. japonicum (top in each figure) and C. magnificum (bottom left in each figure).* ITS haplotype map and haplotype network of Platycrater arguta.
|
GSK3 gene family evolutionGene fate after duplication is still a mystery to us; the evolutionary history of large gene families is usually extremely complex. The increasing availability of genome and transcriptome data allows us to including almost entire gene family members for large gene family. Benefitting from the 1KP project and Amborella genome project, we are able to reconstructed better phylogenies for some large gene families across land plants.
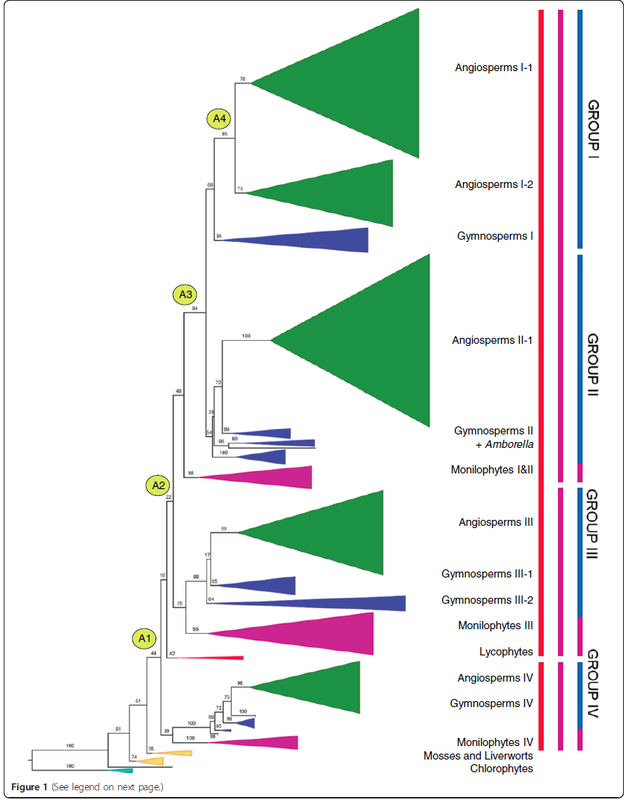
GSK3 genes encode signal transduction proteins with roles in a variety of biological processes in eukaryotes. In contrast to their low copy numbers in animals, GSK3 genes are numerous in land plants and have diverse functions, including floral development in angiosperms. Five GSK3 loci that were present in the ancestral angiosperm have subsequently diversified among major angiosperm lineages, but a sixth ancestral locus has been detected only in Amborella. Thus, among flowering plants, Amborella alone may contain all the GSK3 gene lineages that arose before the origin of extant angiosperms, underscoring the importance of Amborella for reconstructing the ancestral angiosperm genome. I performed this work with Dr. Andre Chanderbali while visiting the Soltis lab at University of Florida. More details about this research is available here and here (Science). |
* Phylogenetic relationships among land plant GSK3 genes
|